Genomics research results in zebrafish research (Genomics: Zebrafish earns its stripes)
Author: Xie Wei Xun
The processes of human development, physiological function and disease involve thousands of genes and their variants, but the function of most genes and their variants remains unknown. In the past 20 years, zebrafish has gradually become an important model animal for studying the function of human genes. In two articles published on Nature's website, the zebrafish reference genome sequence and the identification of rupture mutants that complete more than 10,000 protein-coding genes have been reported, which, together with recent research reports, will strongly promote human disease research.
A common means of studying gene function is to determine the effects of a particular mutation on the phenotype of the body, such as bones, physiology, and behavioral patterns. Zebrafish embryos and juveniles have the characteristics of small size, easy access, and transparent embryo body. They can perform a large number of in vivo detection and analysis at the single cell level, which is very suitable for this research. Most of the zebrafish gene function is revealed by "forward genetics", which first introduces random mutations in the zebrafish genome and identifies phenotypic changes in the progeny (fig. 1a). The use of this method to identify teratogenic mutations is very labor intensive, but to a large extent helps people to reveal many genetic pathways from embryonic development to cardiac physiology regulation.
The high-quality zebrafish genome sequence reported by researchers by Howe et al. makes it possible to directly align the mutated sequence with the wild sequence, which will greatly facilitate the identification of the mutation. The zebrafish genome sequence shows that 75% of human disease-related genes can find homologous genes in zebrafish, so we can use zebrafish for functional studies of disease-related genes.
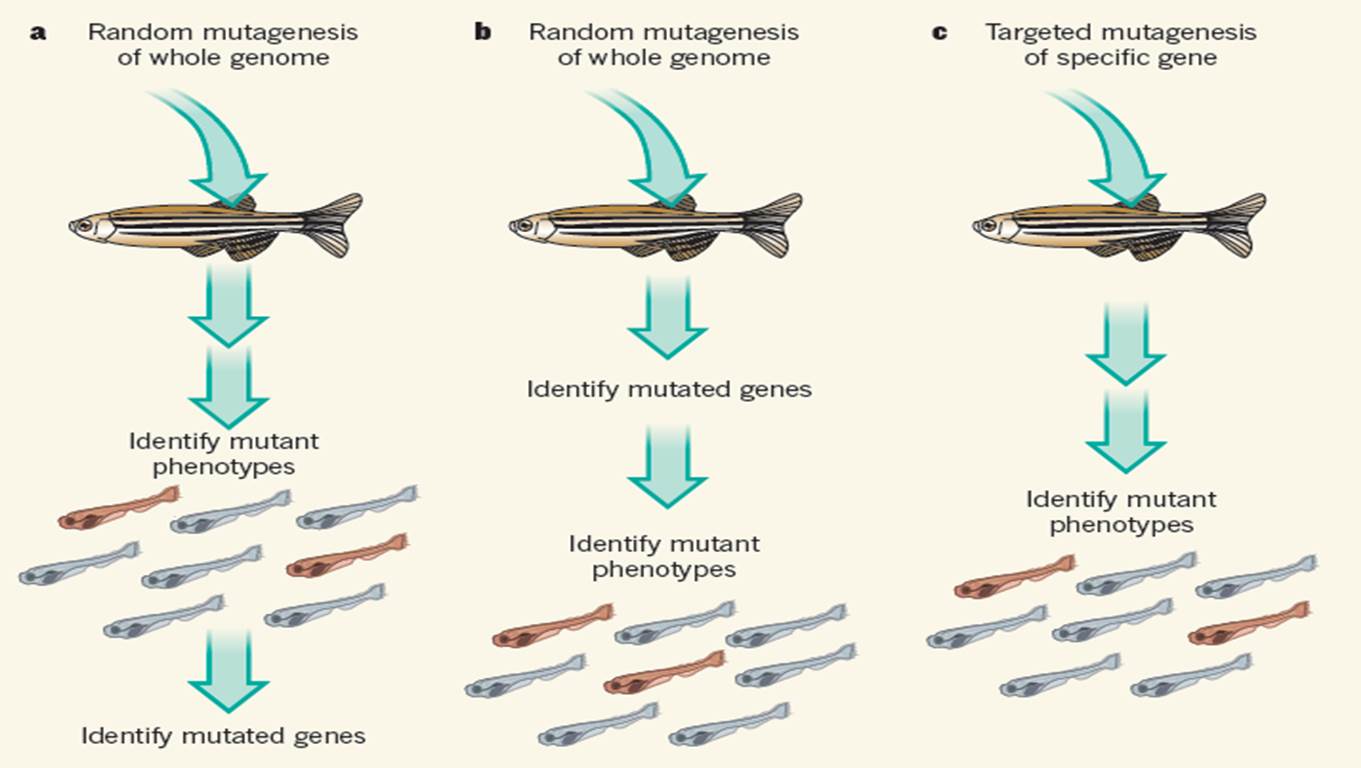
Figure 1 shows the relationship between phenotype and genes . Three methods commonly used in zebrafish to study the function of specific genes in the body's phenotype. a , Forward genetics, introducing random mutations into adult fish, identifying phenotypic variations in the offspring, and analyzing the genome for mutations. b . Another method is to perform a random mutation first, but then compare the progeny genome sequences to find mutations to determine the phenotypic changes. c . A mutation technique for a specific target gene, introducing a mutation to a specific gene and then analyzing the phenotype of the progeny.
But how do you study the function of a particular human gene in zebrafish? First, mutations need to be introduced into the zebrafish homologous gene, followed by phenotypic analysis (fig. 1b). Kettleborough et al confirmed that similar experiments can be performed on a large scale. The authors randomly mutated the male zebrafish and subsequently sequenced the protein coding regions of the progeny genomic DNA. In 1,673 fish, the researchers identified 10,043 genes (more than one-third of all zebrafish protein-coding genes) were disrupted. These mutant lines will be a valuable resource for systematic research on the function of these genes.
Another way to study gene function is to introduce mutations into specific genes rather than introducing random mutations to the entire genome (fig. 1c). With the emergence and improvement of techniques for shearing DNA at specific sites, genetic mutation techniques have undergone fundamental changes. The latest reported technology is the CRISPR-Cas9 system, which has been successfully applied to zebrafish. At the heart of the technology is an RNA molecule that is complementary to a partial sequence of the target gene, directs the endonuclease to a specific DNA site of the target gene, cleaves it, and introduces the mutation through an error-prone repair mechanism. The relatively large-scale mutation screening of the CRISPR-Cas9 system is cheaper, faster, and can be used by smaller laboratories. Therefore, the production of corresponding mutants of most zebrafish genes is only a matter of time.
Despite these breakthrough discoveries, it is not clear how many genes will produce corresponding phenotypes in the event of damage. Based on previous positive genetic screening results, it is speculated that less than 10% of the zebrafish gene disruption will produce an abnormal phenotype during the first 5 days of embryonic and juvenile fish. Consistent with this, Kettleborough et al found that only about 5% of the more than 800 genes were required for normal development during this period.
Does this mean that more than 90% of the zebrafish genes are functionally unrelated? There are a few considerations below that make this statement completely untenable. First, Kettleborough and colleagues included only very obvious structural features for phenotypic analysis, and relatively subtle phenotypes were often overlooked. Second, many genes only function in late development or adulthood. Third, in the early developmental stages, disruption of gene function is often compensated by maternal factors present in the yolk or embryo body. Finally, genes with similar sequences often have overlapping or partially redundant functions, resulting in the destruction of a single gene without a phenotype or a slight phenotype.
Howe et al found that a quarter of the zebrafish genes have homologous genes with high sequence similarity, suggesting that the last point above is particularly useful in zebrafish. However, all of these possibilities can be solved by sophisticated techniques that can be applied to zebrafish: phenotypic analysis can use high-resolution imaging techniques and gene expression profiling; mutant fish can be used for late developmental studies; It is relatively easy to knock out 2-3 related genes at the same time as the mutant fish expressed in the zygote phase.
How will these tools and resources accelerate the study of human disease-related genes? The method is well understood, looking for homologous genes in human zebrafish genes and editing them, analyzing abnormal phenotypes, using high-throughput drug screening platforms to find or identify small molecule compounds that can modulate this phenotypic change. In addition, the completion of the zebrafish genome sequence allows us to use new research methods to solve problems that have long plagued humans. For example, zebrafish have a high proportion of genetic variation - Howe et al found that there is a variation every 200 bp in different strains and even in different individuals. Studies in other organisms have shown that such differences can lead to phenotypic effects. It is conceivable that zebrafish will be a useful spinal model animal for studying the role of minor genotypic variation in phenotypic differences.
Will the completion of the zebrafish genome sequence introduce new ideas in biology? very possible. It is clear that the publication of the human genome sequence has not brought about major changes. For example, no one predicts the important role of a large number of non-coding protein RNA molecules in functional regulation, or is it foreseen to use genomic sequences to reconstruct human evolutionary history? The collection of zebrafish genome sequences and mutants may be a precursor to the next major discovery .
Alexander F. Schier is currently working at Harvard University's Department of Molecular and Cytology, Cambridge, Massachusetts 02138 , USA
Email: vard.edu
1 Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature, doi:10.1038/nature12111 (2013).
2 Kettleborough, RN et al. A systematic genome-wide analysis of zebrafish protein-coding gene function. Nature, doi:10.1038/nature11992 (2013).
3 Varshney, GK et al. A large-scale zebrafish gene knockout resource for the genome-wide study of gene function. Genome research 23 , 727-735, doi:10.1101/gr.151464.112 (2013).
4 Hwang, WY et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nature biotechnology 31 , 227-229, doi:10.1038/nbt.2501 (2013).
5 Chang, N. et al. Genome editing with RNA-guided Cas9 nuclease in Zebrafish embryos. Cell research 23 , 465-472, doi:10.1038/cr.2013.45 (2013).
6 Zu, Y. et al. TALEN-mediated precise genome modification by homologous recombination in zebrafish. Nature methods 10 , 329-331, doi:10.1038/nmeth.2374 (2013).
7 Bedell, VM et al. In vivo genome editing using a high-efficiency TALEN system. Nature 491 , 114-118, doi:10.1038/nature11537 (2012).
8 Driever, W. et al. A genetic screen for mutations affecting embryogenesis in zebrafish. Development 123 , 37-46 (1996).
9 Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123 , 1-36 (1996).
10 Solnica-Krezel, L., Schier, AF & Driever, W. Efficient recovery of ENU-induced mutations from the zebrafish germline. Genetics 136 , 1401-1420 (1994).
11 Zon, LI & Peterson, RT In vivo drug discovery in the zebrafish. Nature reviews. Drug discovery 4 , 35-44, doi:10.1038/nrd1606 (2005).
12 Lander, ES Initial impact of the sequencing of the human genome. Nature 470 , 187-197, doi:10.1038/nature09792 (2011).
Intermediate for CAS 175591-09-0
Tapentadol Hydrochloride Intermediate,Tapentadol Hcl,Tapentadol Hcl Api,Tapentadol Intermediates
Shandong Bolode Bio-technology Co., Ltd. , https://www.bldpharma.com