Psoriasis, commonly known as cognac, is a common chronic inflammatory skin disease with characteristic skin lesions and is considered to be a polygenic genetic disease in which genetic factors interact with various factors such as environmental factors.
In the past, the pathogenesis of polygenic genetic diseases was explored. The researchers found a correlation between polymorphisms in the two groups of individuals and the genetic disease through a SNP-based genome-wide GWAS experiment of normal and disease control samples. However, the onset of many diseases is acquired. For example, diabetes is closely related to the diet, that is, the acquired living environment. The genetic mechanism of such complex diseases is difficult to find all the decisive genetic mutations through GWAS based on SNP chips. With the improvement of second-generation sequencing technology and sequence enrichment technology, people have turned their attention to the disease-related gene coding region.
Whole-genome exon sequencing is a genomic analysis method that uses sequence capture technology to capture and enrich the whole genome exon region DNA for high-throughput sequencing. Because of its high sensitivity to common and rare mutations, it can discover the vast majority of disease-related variability in the exon and the need to sequence only about 1% of the genome, enabling the whole genome exon to be sequenced and based on its development. Targeted capture technology is the most effective strategy for identifying pathogenic genes for Mendelian disease, and is also used in the research and clinical research of complex disease susceptibility genes. 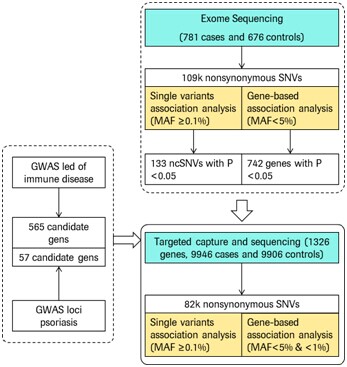
Recently, at the First Affiliated Hospital of Anhui Medical University, he led a number of research institutions involved in psoriasis project jointly published an article in Nature Genetics 1, combined exome capture and targeting region and second-generation Sequencing technology provides a new technical route for the study of the mechanism of psoriasis, a polygenic genetic disease (see image to the right).
To examine the effects of functional coding region mutations on the genetic susceptibility to psoriasis, Chinese researchers performed a large range of functional region mutations on two completely independent experimental groups (a total of 21,309 individuals, all from the Han population). Sequencing and analysis of the spots, and found six genes associated with the disease and a number of mutations that may cause disease.
In the early stage of the experiment, the researchers used the NimbleGen full exome capture technique to capture the exon regions of all genes from 781 psoriasis patients and 676 normal individual DNA samples, and detected them by sequencing analysis and single mutation correlation analysis. There were 133 non-synonymous mutant SNVs that were significantly associated with the genetic susceptibility to psoriasis and 742 genes that may be associated with disease. However, Bonferroni correction results showed that these 742 genes were not significant at the genome-wide level.
Taking into account the effect of sample size on the above experimental results, the researchers used targeted capture sequencing technology to further screen and validate the above mutation sites and pathogenic genes in 9,946 psoriasis patients and 9,906 normal individuals. They used the NimbleGen sequence to capture a custom pair of exons containing 1,326 genes covering 133 non-synonymous mutant SNVs, 742 possible causative genes, and 622 genes that have been detected by GWAS as immune disease-related genes. After the region is enriched, deep sequencing is performed again.
Combining the results of a single mutation correlation analysis of non-synonymous mutant SNVs from 10,727 cases of two experimental populations and 622 immune disease-related genes from 10,582 controls, the researchers identified six possible psoriasis-causing genes, namely : IL23R, LCE3D, ERAP, GJB2, ZNF816A and FUT2.
Based on the results of the above experiments, the researchers indicated that more positive results were obtained in the validation of subsequent populations in the susceptibility genes of previous GWAS studies by targeted enrichment deep sequencing; for genetic diseases, especially polygenic genetic diseases The pathogenesis research, this new technical research route, the whole exome and small-scale study of targeted region enrichment, can deepen the sequencing depth while focusing on the coding of functional genes without generating a large amount of sequencing costs. The search for mutation-related mutation sites in the region provides a strong complement to previous SNP-based GWAS studies and provides more biomarkers for future clinical diagnosis.
1. A large-scale screen for coding variants predisposing topsoriasis. Nature Genetics. 2014 Jan. DOI:10.1038/ng.2827
Butyl Rubber Stopper For Prefilled Syringes
Our rubber stoppers are made of chlorobutyl rubber materials which obeys the FDA rules , and our products are widely used in all kinds of domestic and imported medical products according to the different needs of our customers .Recently our 10mm Medical Chlorobutyl Rubber Stoppers are on hot sale. it can be used in high therapeutical infusion products, sensitive powder injection medicine,blood plasma products and so on. Moreover, the products have good sealing and low water-gas transmittance, good compatibility with drugs. It has a low dissolution when our product is tested in lab which due to its pure formula,sulfur-free, zinc-free vulcanization system. It also has good puncture retention and low chip rate. If you are looking for a Butyl Rubber Stopper supplier in china , we believe we are your best choice. Looking forward to your inquiries.
Butyl Rubber Stopper For Prefilled Syringes,Stopper For Prefilled Syringes,Rubber Stopper For Injection,Round Butyl Rubber Stopper
SUZHOU CRH PHARMACEUTICAL TECHNOLOGY CO.,LTD , https://www.crh-health.com
