The perfect combination of impurity protein identification and quantification - Hi3 analysis method
Jia Wei
Waters Technology (Shanghai) Co., Ltd. Experimental Center
In the liquid quality data, not only the peptide map information containing the correct expression of the protein drug sequence, but also other impurity proteins can be excavated. However, the identification of pure impurity proteins is not far enough, and the exact content information is equally important. Waters PLGS software can further identify the impurity proteins in the peptide map liquid data, and analyze the relative content information.
I. Identification and relative quantification of impurity proteins completed by "by the way"
Using the corresponding open database, the PLGS software first identifies the peptide based on the ion fragmentation information in the liquid data, and then assembles all the proteins in the sample to complete the protein identification step. In the relative determination of protein content, PLGS software makes full use of the data advantages of the MS E full information tandem mass spectrometry method. Compared with the DDA method, the ion chromatographic peak of MS E data is almost “perfectâ€, which is more in line with the actual peptide peak shape. Through comprehensive research, it was found that the peak intensity information of the peptide in the MS E data is very correlated with the protein concentration. Each protein was used to identify the sum of the peak intensity of the top three polypeptides in the polypeptide, ie, to characterize its protein concentration. Therefore, using PLGS to analyze the same peptide map liquid data, you can "by the way" to complete the identification and relative quantification of impurity proteins.
Second, the determination of absolute protein content
It is also very simple to get accurate protein absolute concentration information in a sample. Simply add a known protein internal standard to the sample. At this time, the Hi3 analysis method quantifies the absolute concentration in the sample according to the concentration of the internal standard protein according to the following formula [1, 2].

3. Analysis of residual host cell protein (HCP) using the Hi3 method
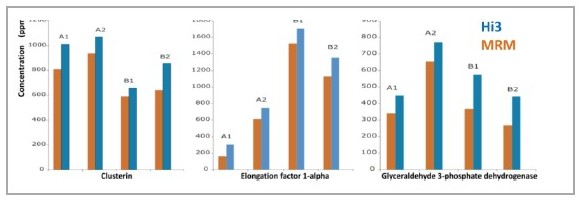
The fusion antibody anti-phosphotyrosine IgG 1 (PTG1 mAb) was expressed in two culture conditions using the Chinese hamster ovary cell line DG-44 CHO, and the expressed PTG1 mAb was purified by A and B methods, respectively. After the liquid chromatography data was collected by the 2D-UPLC MSE method, it was analyzed using PLGS software [3]. In the analysis results, the impurity proteins contained in the sample can be clearly given, and the relative content of these impurity proteins can be given (Fig. 1). In order to verify the reliability of the Hi3 quantitative method, the MRM method was used to quantitatively verify the three proteins of Clusterin, Elongation factor 1-alpha and Glyceraldehyde 3-phosphate dehydrogenase. By comparing the results of the two quantitative methods of Hi3 and MRM, it can be found that the quantitative concentrations of the two are very close (Fig. 2). In the identification and quantification of impurity proteins in samples using PLGS, the search method setting is almost identical to the peptide map analysis, and only the step of adding the name and concentration of the reference protein to the analysis parameter is added. After that, all the analysis will be done by PLGS and a list of results for protein identification and concentration will be given directly. In fact, PLGS is not only used in biopharmaceutical impurity protein analysis, in the extremely complex proteomics study of samples, Hi3
Methods have been used for a long time, and many high-level papers have been published [4, 5, 6].
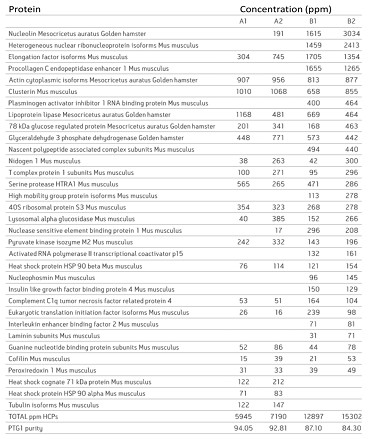
Figure 1. Identification results and content of impurity proteins.
references
(1) Silva JC, Gorenstein MV, Li GZ, Vissers JP, Geromanos SJ. Absolute Quantification of Proteins by LCMS. Mol Cell Proteomics.2006, 5, 144-156
(2) Silva JC, Denny R, Dorsc hel C, Gorenstein MV, Li GZ, Richardson K, Wall D, Geromanos SJ. Simultaneous Qualitative and Quantitative Analysis of the Esc heric hia coli Proteome. Mol Cell Proteomics. 2006, 5, 589 -607.
(3) Catalin Doneanu, Keit h Fadgen, St Jo hn Skilton, Mart ha Sta pel s, Weibin C hen. A generic 2D-UP LC/MS assay for the identification and quantification of host cell proteins in biopharmaceuticals. Waters application note 2011 , 720004043en
(4) Tenzer S, Wee E, Burgevin A, Stewart-Jones G, Friis L, Lamberth K, Chang CH, Harndahl M, Weimershaus M, Gerstoft J, Akkad N, Klenerman P, Fugger L, Jones EY, McMichael AJ, Buus S, Schild H, van Endert P, Iversen AK. Antigen processing influences HIV-specific cytotoxic T lymphocyte immunodominance. Nat Immunol. 2009, 10,
636-646
(5) Stapels M, Piper C, Yang T, Li M, Stowell C, Xiong ZG, Saugstad J, Simon RP, Geromanos S, Langridge J, Lan JQ, Zhou A. Polycomb
Group proteins as epigenetic mediators of neuroprotection in ischemic tolerance. Sci Signal. 2010, 3, 111, ra15
(6) Wang X, Kota U, He K, Blackburn K, Li J, Goshe MB, Huber SC, Clouse SD. Sequential transphosphorylation of the BRI1/BAK1 receptor
Kinase complex impacts early events in brassinosteroid signaling. Dev Cell. 2008, 15, 220-35.
FOLDUP AND COMPACT: Providing a convenient option for those needing assistance, the Rohi folding stick smoothly and securely locks in place to provide extra stability while walking on any surface. Folding at three points along the shaft, the folding stick is compact, easily fitting into a purse, briefcase or wheelchair bag.
Folding Adjustable Walking Stick/Cane ,Walking Stick Cane for Men Women,Portable Cane with T Handle
jinan Fangqi Sunshine education consulting CO.,LTD. , https://www.fzrehabmarket.com